Biology is nothing short of alchemy—the transmutation of matter and energy into precious materials. It is well known that the genome of each organism encodes the ‘hardware,’ or physical set of enzymatic machinery, which enable these transformations. By becoming proficient in this genetic language, we may one day be able to unlock a future of distributed manufacturing which partners with living systems at the basis of our economies. To bring this vision forward, I spent the Summer of 2021 building OpenAlchemy, an open-source software that automates DNA design for molecular manipulation.
Database
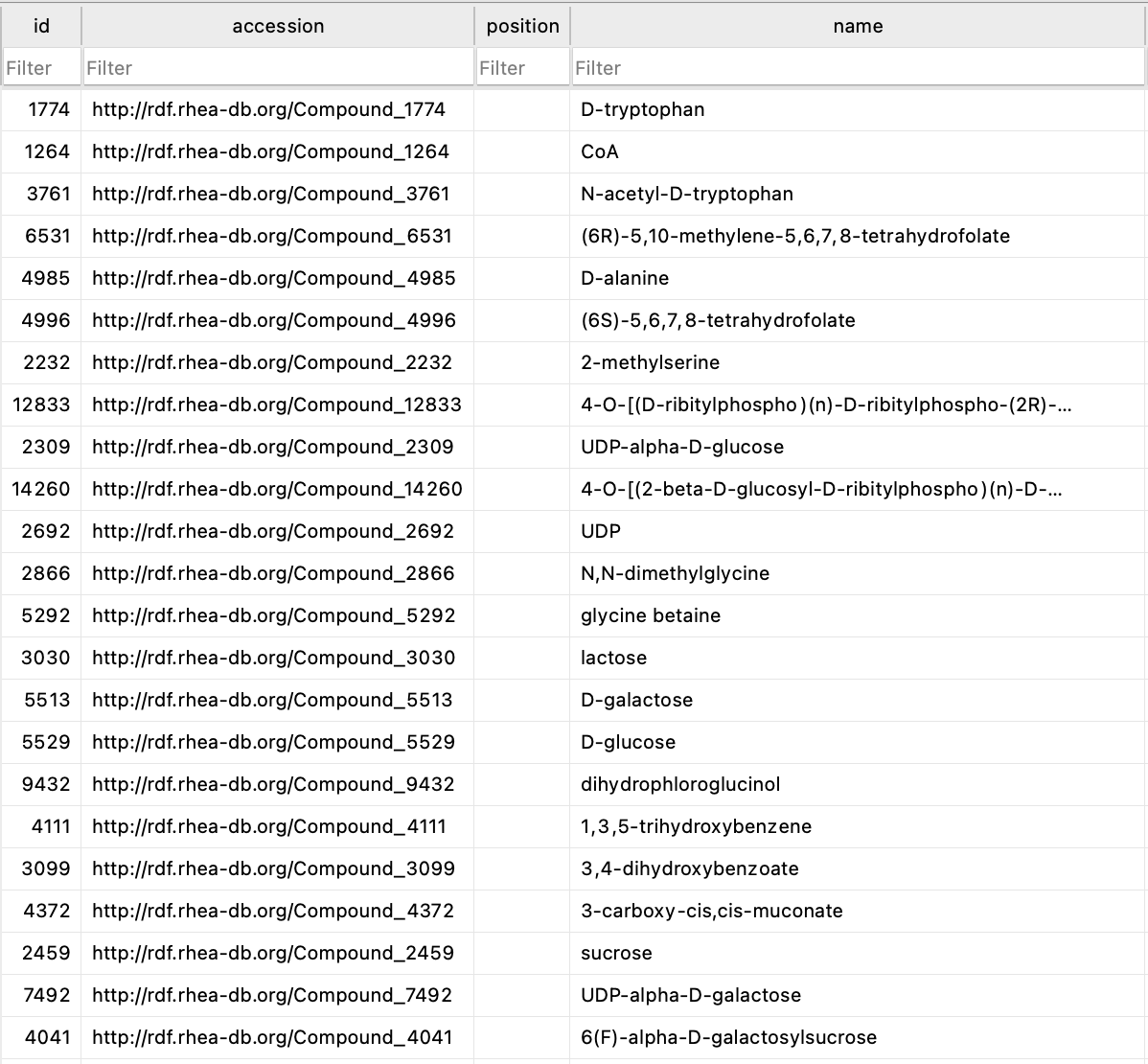
Working with fellow bioengineer Keoni Gandall, I compiled all the data from Genbank (genes), Uniprot (proteins), and Rhea (enzyme reactions) into one database.
SQL
With this data centralized, I taught myself SQL and created queries that start with an end molecule in mind, and recursively trace through the entire database until a pathway ends in a substrate-product pair that is contained within the chassis organism to be engineered.
Golang
Finally, I taught myself Go and developed methods to integrate the backend querying with a command-line user interface.
Output
As an example output, here are the required DNA sequences that are required to convert xanthosine monophosphate into ATP.
For more details, please see the full code documentation at the Github link.
Future Directions
Currently, I’m working to redevelop this with a simpler UI/UX in Python and extend functionality to automated plasmid design, metabolic pathway optimization and ML-powered de novo enzyme generation.